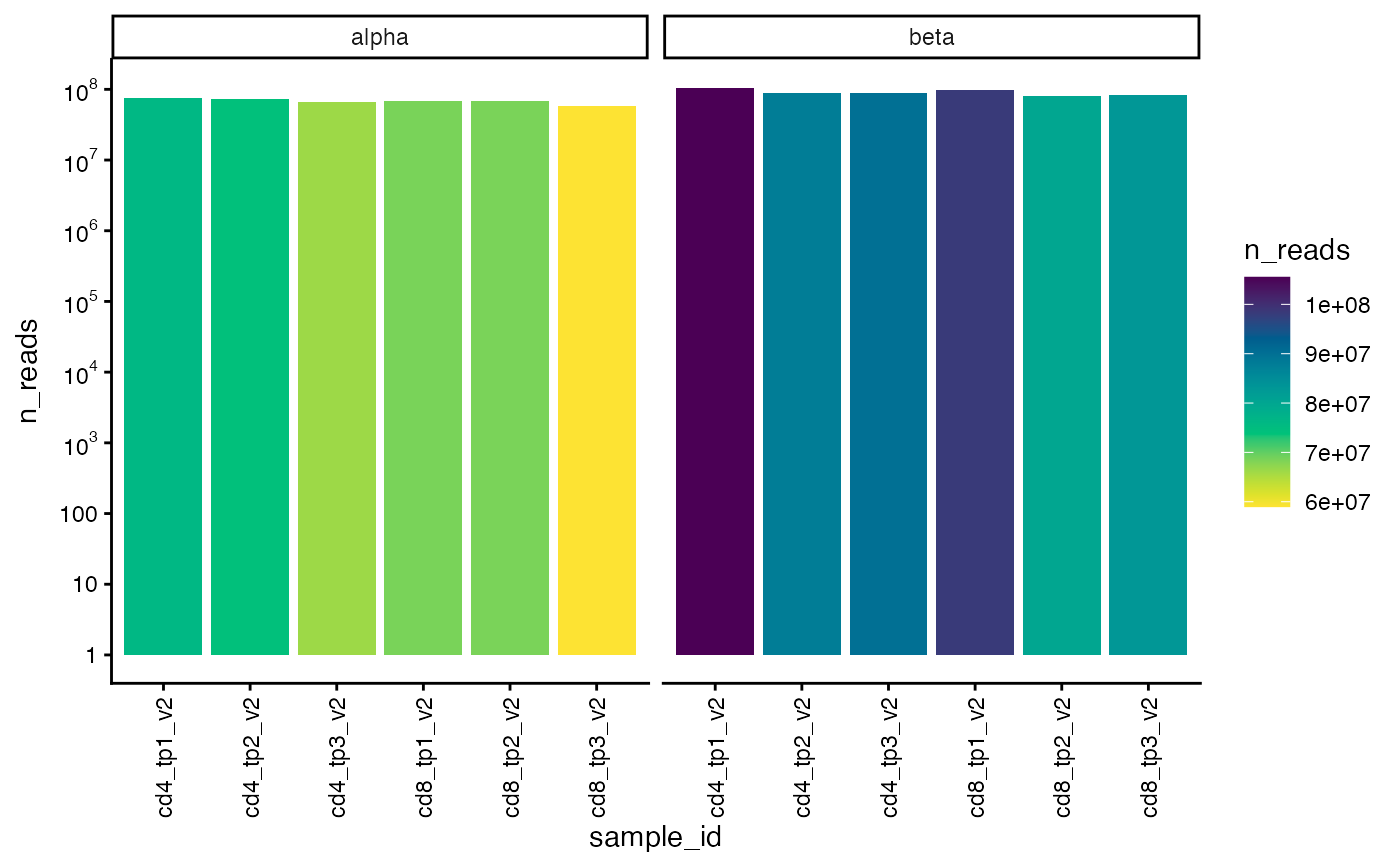
![[Experimental]](figures/lifecycle-experimental.svg) This function plots the total number of alpha and/or beta reads (default is both)
for each sample.
This function plots the total number of alpha and/or beta reads (default is both)
for each sample.
Usage
plot_n_reads(
data,
chain = c("both", "beta", "alpha"),
samples = NULL,
color_scheme = NULL
)
Arguments
- data
a TIRTLseqDataSet object
- chain
the TCR chain to plot (default is both "alpha" and "beta")
- samples
(optional) which samples to plot (default is all samples)
- color_scheme
a color scheme for the plot
See also
Other qc:
get_all_tcrs(),
get_pair_stats(),
get_paired_by_read_fraction_range(),
plot_num_partners(),
plot_paired(),
plot_paired_by_read_fraction_range(),
plot_paired_vs_rank(),
plot_pairs_with_eachother(),
plot_ranks(),
plot_read_fraction_vs_pair_status(),
plot_sample_overlap(),
summarize_data()
Examples
folder = system.file("extdata/SJTRC_TIRTL_seq_longitudinal", package = "TIRTLtools")
ts_data = load_tirtlseq(folder, meta_columns = c("marker", "timepoint", "version"), sep = "_", verbose = FALSE)
plot_n_reads(ts_data)